Submitted by Sushma_Naithani on Wed, 05/07/2025 - 13:27
The April 2025 public release (Version 24) of the Plant Reactome pathway knowledgebase hosts pathways for 139 species representing model plants, crops, and species of evolutionary importance. It is an open-source and freely accessible resource, built upon the Neo4j graph database management system to facilitate a seamless integration of heterogeneous data
Submitted by Sushma_Naithani on Fri, 05/21/2021 - 18:51
Plant Reactome is the pathway portal of Gramene knowledgebase.We combine manual biocuration and gene-orthology based automated projections to provide conceptual framework of system-level plant pathway networks for 106 species of photoautotrophs, including models, crops and species of evolutionary interests.
Submitted by Sushma_Naithani on Tue, 05/26/2020 - 14:25
The advent of functional genomics has changed the scope, scale and speed at which new datasets are being generated and has posed the challenge for its comprehension and effective utilization to advance translational research. Plant Reactome portal of Gramene
Image:
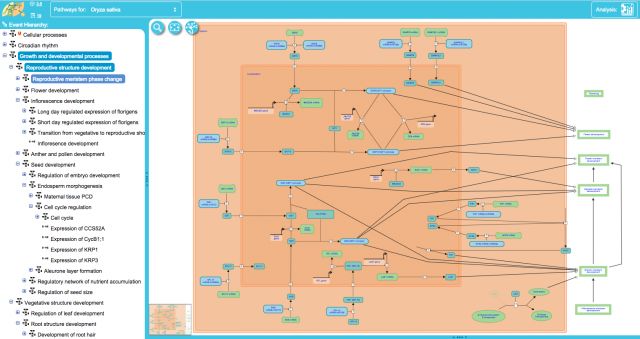
Reproductive meristem phase change in O. sativa
Submitted by Sushma_Naithani on Wed, 05/13/2020 - 12:21
We wish and hope that in this difficult time of COVID-19 pandemic, all Gramene database users and their loved ones are doing well and keeping safe.
Submitted by Sushma_Naithani on Wed, 01/22/2020 - 12:31
The recent 2020 Plant and Animal Genome (PAG) Conference, held in Town & Country Resort, San Diego, brought together over 3,000 leading researchers, scientists, software professional, and students working in plant and animal genomics research, over 135 exhibits, 180 workshops, 1,200 posters, over 2,500 abstracts and 7 Plenary talks.
Submitted by Sushma_Naithani on Thu, 01/09/2020 - 17:35
The Plant and Animal Genome (PAG) XXVIII conference will kick off this coming Saturday, January 11th, 2020, in San Diego, CA. Come meet Gramene staff and get the latest updates to Gramene’s comparative genomics and pathways visualization and mining tools to aid your research on plant models and crops! At PAG 2020, we will be announcing Gramene's p
Submitted by Sushma_Naithani on Tue, 11/12/2019 - 13:50
The Gramene is a curated, open-source, integrated data resource for comparative functional genomics in crops and model plant species.
Submitted by Sushma_Naithani on Fri, 02/08/2019 - 13:56
The Gramene Team is pleased to announce its Release #60 with the Genome section providing access to information on 2,162,056 genes and 58 reference plant genomes. 1,891,391 protein-coding genes are organized in 93,194 gene families.
Submitted by Sushma_Naithani on Tue, 01/08/2019 - 14:33
The Gramene team looks forward to meeting plant researchers at the Plant and Animal Genome XXVII conference from 12th-16th January, 2019. You can meet Gramene folks at:
Submitted by Sushma_Naithani on Thu, 08/23/2018 - 14:17
Image:
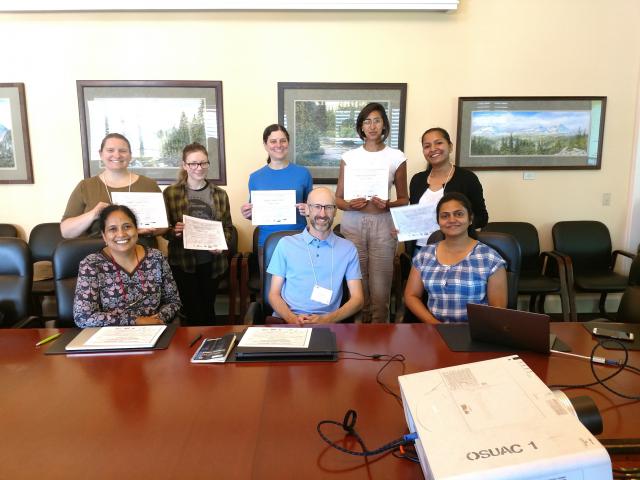

Photo: Left to right in the front row are Sushma Naithani, Justin Preece, Parul Gupta and in the back row are Valerie Fraser, Lillian K Padgitt-Cobb, Kelly Vining, Noor-Al-Bader and Priyanka Garg.
Pages